In the introduction to data science article, we laid the foundation for data science projects by explaining how to set up a data science team and the objectives that need to be met. We noted the data scientist is responsible for analyzing the data to provide insights to decision makers. There are multiple tools available such as Python and R. In this article, we will demonstrate how to set up an R environment, import data into R from different sources and explore the data to understand it. In this article, a basic understanding of R is assumed.
RStudio is an integrated development environment (IDE) that helps data scientists improve their productivity. RStudio provides features for syntax management such as highlighting, code execution, and code completion. Another benefit of RStudio is that it brings together the different tools that are needed in a data science workflow. Both commercial and open source versions of RStudio are available for Windows, Mac and Linux desktops. Server versions that allow you to access Rstudio via a browser are available for Linux distributions such as Ubuntu, Suse, RedHat and CentOS.
To set up a desktop development environment, you need to begin by downloading and installing the latest R version on your Windows, Mac or Linux desktop. R is available from the CRAN site at https://cran.r-project.org/. After installing R, the next step is downloading and installing RStudio from here https://www.rstudio.com/products/rstudio/. At this stage a development environment is completely set up and we can begin importing data into R. The screenshot below shows a correctly set up development environment.
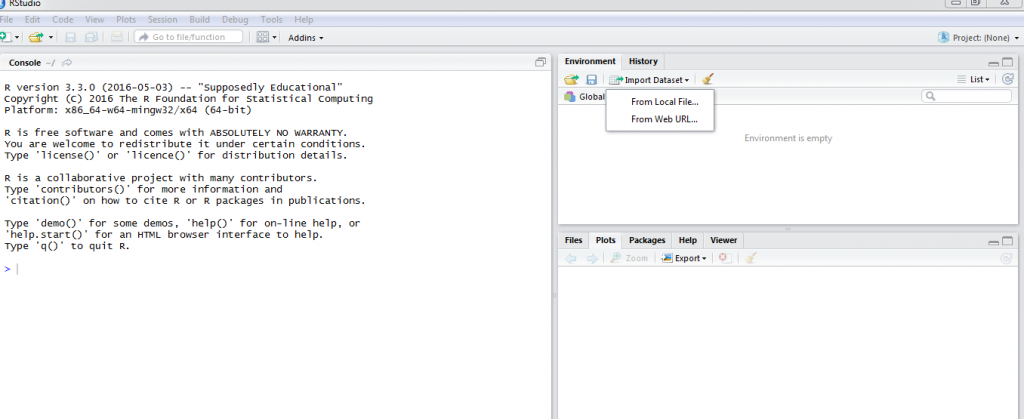
When using Rstudio, Knitr provides a very efficient way of documenting your code and the output from your code. With Knitr you are able to produce HTML, PDF or Microsoft Word documents that contain either the code, output or both. To install the knitr package execute this command install.packages(“knitr”) at a the R console.
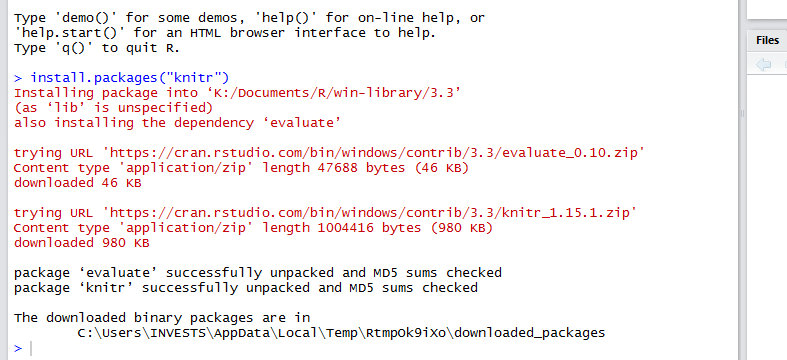
After installing knitr, we are ready to start documenting our code using HTML documents. Click on file, new file then R Markdown. You will be prompted to select the type of document and specify a title for your document.
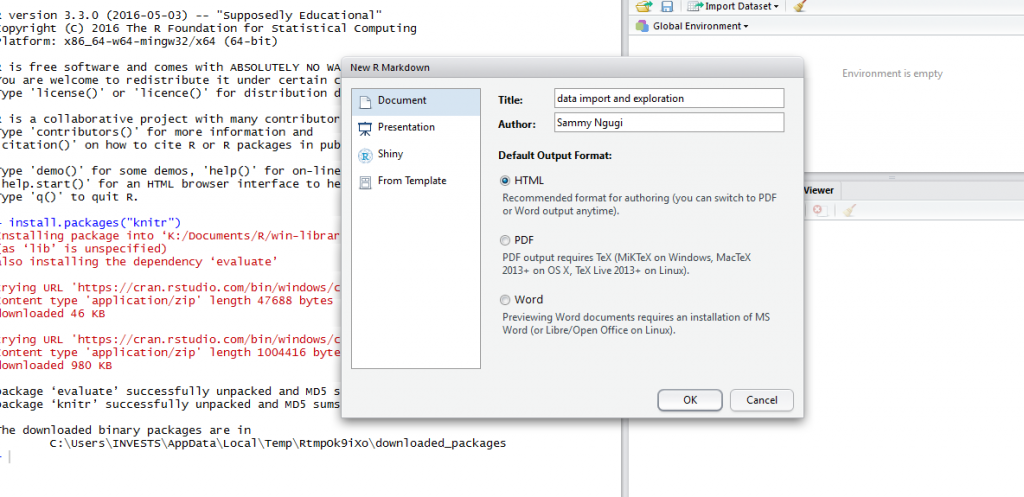
After clicking OK, a new R Markdown document will be created as shown below.
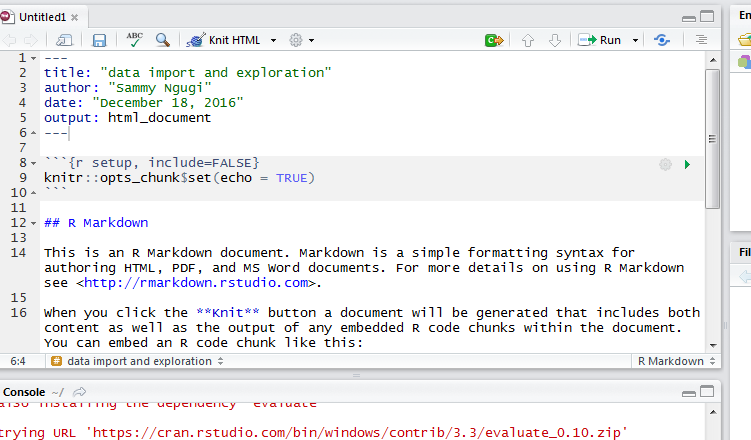
In an R Markdown document, you place your chunk of R code within the three opening and closing tildes i.e. the three tick like symbols. You can create multiple code chunks. There are several options that are used to control how your document will be created. In the first line after the opening tilde symbols you specify you are writing R code, you specify the name of the code chunk and specify options. The echo = FALSE option is used to hide the code while showing the results. The results=”hide” is used to show the code but not the results. The include=FALSE option is used to evaluate the code but hide the output and the code. These are just examples of the options that can be used, an exhaustive list of options is available here http://yihui.name/knitr/options/#chunk_options.
R is able to consume different data sources. The most common data source are well structured files that have data arranged in rows and columns. Whenever you are faced with the problem of manipulating data files to fit into a required structure it is better to write R scripts instead of manually formatting your data. Common data formats are text and CSV that can be delimited using tabs, commas or space. The files can also have column headings in the first line of data or not. When importing your data you need to specify all of these options. For example download the zipped data from this link https://www.fueleconomy.gov/feg/epadata/vehicles.csv.zip and extract it into a local directory. The data has the first line as the column names and it is a CSV file so we need to pass this information when importing the data. The command below imports the data into R.
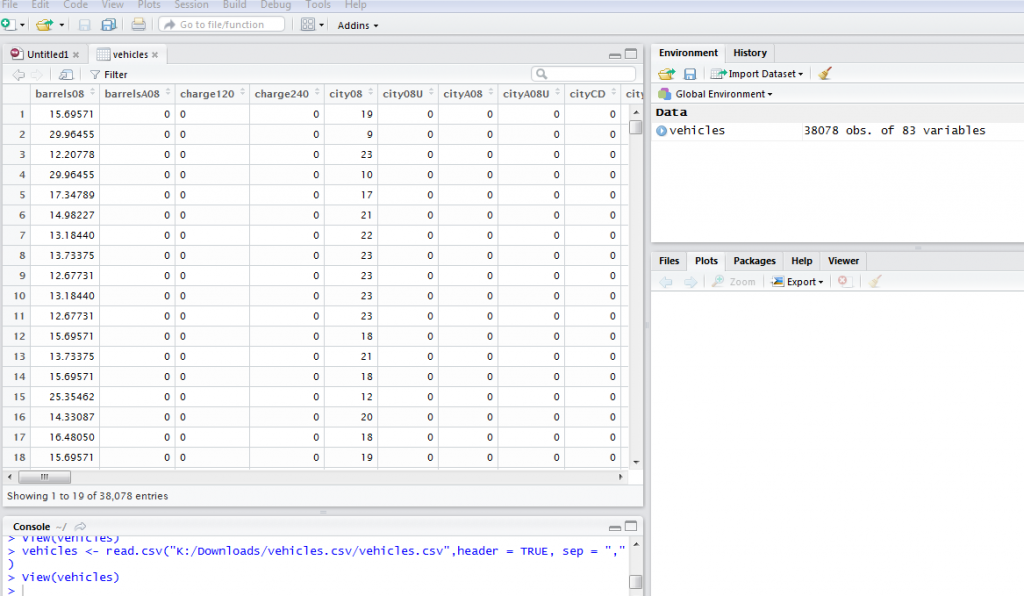
vehicles <- read.csv("K:/Downloads/vehicles.csv/vehicles.csv",header = TRUE, sep = ",")
The data is imported and we can use the command view(vehicles) to display the data in a spreadsheet style.
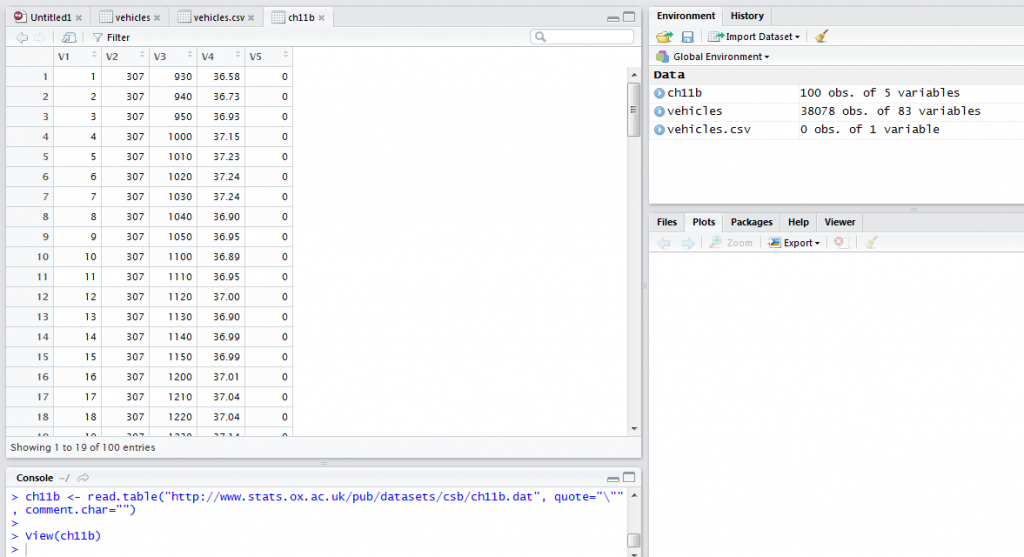
To load data that is well structured from a URL we need to specify the URL, if the first line has column names and the type of separator. For example, to load data from this URL http://www.stats.ox.ac.uk/pub/datasets/csb/ch11b.dat we would use the command below.
ch11b <- read.table("http://www.stats.ox.ac.uk/pub/datasets/csb/ch11b.dat", quote="\"", comment.char="")
View(ch11b)
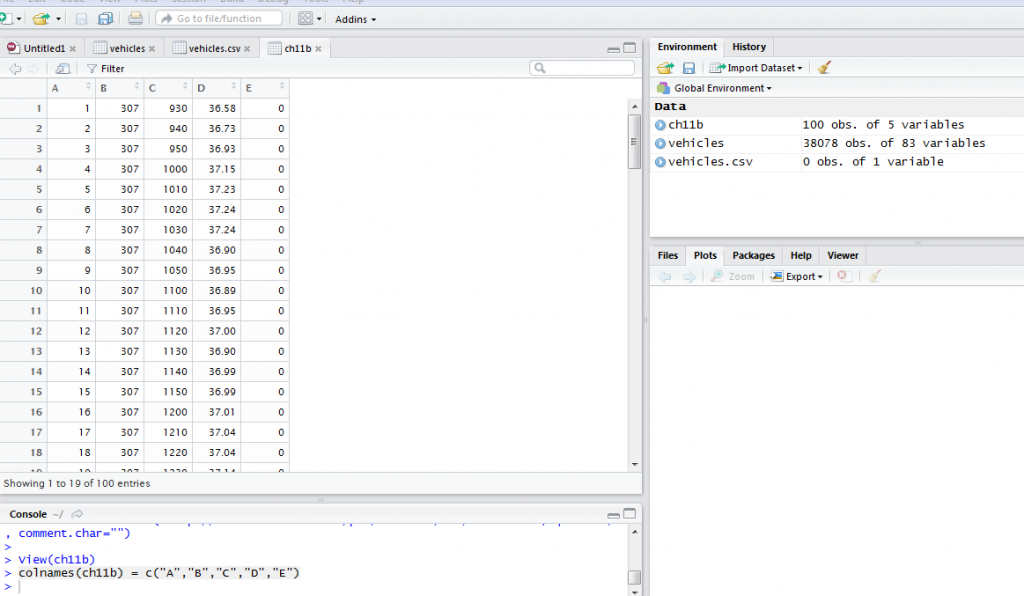
The data we imported from the URL did not have column names. This is an example of some of the data challenges you are likely to encounter. To add column names, we use the command shown below:
colnames(ch11b) = c("A","B","C","D","E")
Databases are very efficient for storing large volumes of structured data. When data we would like to analyze is stored in a relational database, there are two ways it can be imported into R. The first way is to export the data into a file and then import it using the techniques we have discussed previously but this is not the most elegant approach. A better approach is to query the data directly from within R.
In this article, we will use MySql to demonstrate how to import data into R. First, we need to install and load the Rmysql package. The commands below install and load the package.
install.packages("RMySQL")
library(RMySQL)
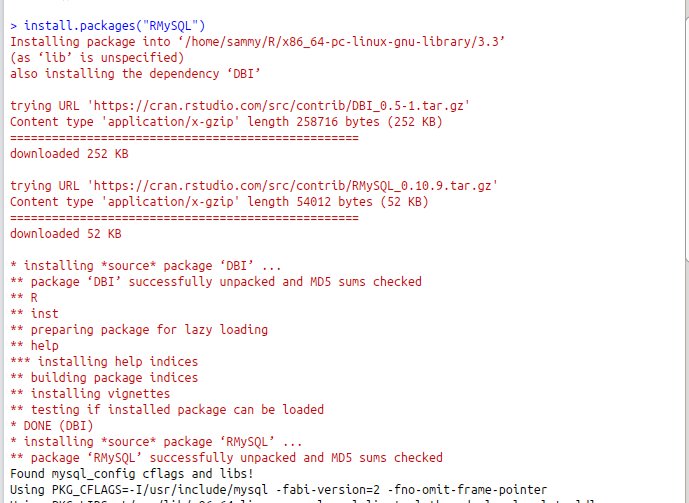
To query data from a MySql database, we need to create a connection object by specifying the database user, password, database name and the host where the database is running. Then, we can run simple or complex queries to bring in the required data.
In this tutorial, we introduced R as one of the important tools to a data scientist. We explained how to set up a development environment using R and Rstudio. We explained some of the features Rstudio provides to improve the productivity of the data scientist. We explained how to use R Markdown to document code and output. We explained how to load local data files and those that reside on the Internet. Finally, we discussed how to load data from relational databases.



After looking over a number of the articles on your web site,
I really like your way of blogging. I book marked it to my bookmark website list and will be checking back soon. Please check out my website as well and let
me know what you think.